 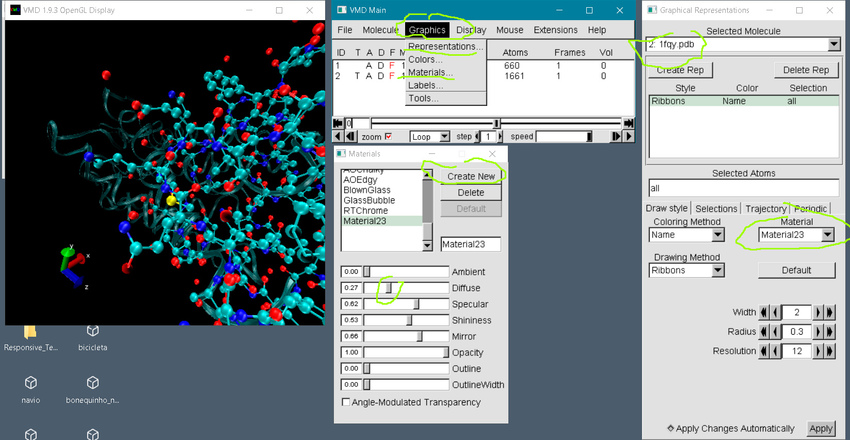 
configure LDFLAGS="-ltcl8.5 -L/mypathtomolfilelibrary/ -L/mypathtotcl" CPPFLAGS="-I/mypathtolibmolfile_plugin. Locate said file and libmolfile_plugin.h, and customize the configure command with something along the lines of. If the init.d folder is not present, just create a new folder and name. If the plugin has a load.tcl file, then go to '\VMD\scripts\init.d' and just paste that load.tcl (or a file named something similar) there. At the end, you should get the molfile plugins compiled as a Copy the plugin folder and go to the path '\VMD\plugins oarch\tcl' and paste the folder there. VMD supports computers running MacOS X, Unix, or Windows, is distributed free of charge, and includes source code. SOURCE of VMD, which contains a plugins directory. VMD is a molecular visualization program for displaying, animating, and analyzing large biomolecular systems using 3-D graphics and built-in scripting. If you configure PLUMED with VMD's plugins you will be able to read 
Could someone give me some tips? Thanks a lot! configure -prefix=/path of plumed/plumed233 LDFLAGS="-ltcl8.5 -L/path/vmd1.9.3/plugins/compile/lib_LINUXAMD64/molfile -L/path/vmd1.9.3/plugins/compile/lib_LINUXAMD64/tcl/" CPPFLAGS="-I/path/vmd1.9.3/plugins/compile/lib_LINUXAMD64/molfile/īut libmolfile_plugin.h was not found during the configuration. I built the plugins successfully (tcl 8.5 was also built), and got libmolfile_plugin.a and libmolfile_plugin.h. cmd file, and modify it.As I want to ask plumed (driver) to process amber NetCDF trajectory files directly, I tried to install PLUMED with vmd plugin.Īs suggested by the Manual, I firstly downloaded the source code of vmd-1.9.3, and there is a plugins directory. mkdir %HOMEPATH%\pymol Then make sure, PyMOL starts here, when you open the shortcut. Now start pymol, and enjoy all the plugins available from the menu. While the computer isn’t powered on yet, press and hold the F2 button of the keyboard, and then press the Power button to enter the BIOS configuration. Firstly, the computer needs to enter BIOS configuration. Then "File->Save as->All files-> C:\Python27\Lib\site-packages\pymol\pymol_path\run_on_startup.py Note: Disable VMD technology will make your computer can’t utilize RAID Array. environ ) # Make setting changes to Plugin Manager import ugins pymol. # Add paths to sys.path so PyMOL can find modules and scripts import sys, os pymol_git = os. Navigate to the installation directory in a CMD window (Not PowerShell!) VMD Store - A VMD extensions that helps users to discover, install. pymol-2.6.0a0-cp311-cp311-win_amd64.whl Visual Molecular Dynamics (VMD) is a molecular modelling and visualization computer program.Download the appropriate wheel files, along with all requirement wheel files ( Numpy+MKL) into a single file directory, e.g., C:\Users\\Downloads.Have a look in "Add/Remove" programs if it's installed already.Systems with equivalent components are also suitable for installation. Otherwise the installed PyMOL binary may fail to run (without any error message!). This package can be installed on systems running the software described below. Install the current Microsoft Visual C++ Redistributable for Visual Studio 2015, 20.Make sure the option to add environment variables is selected or add the folder of python.exe to system PATH. The Force Field Toolkit (ffTK) plugin provides a comprehensive toolset for the development of CHARMM-compatible (e.g. Use the standard options, which should mean that the installation directory is most likely C:\Users\\AppData\Local\Programs\Python\Python38). Install the latest version of Python 3 for Windows (e.g., by going to and choosing the 圆4 EXE installer).Pre-compiled Open-Source PyMOL is available free from Christoph Gohlke of the Laboratory for Fluorescence Dynamics, University of California, Irvine. It also allows sponsors to create highly customized PyMOL installations which might not be possible with the MSI installer. Open-Source PyMOL is available free of charge. The bundle also includes ready-to-use APBS, RigiMOL, an MPEG encoder for movie export, and a small molecule energy minimization engine. Source Code LINUX64 OpenGL, CUDA, OptiX, OSPRay (Linux (RHEL 6. Schrödinger provides an installer to paying sponsors (EXE for PyMOL 2.0, MSI for previous version). We recommend that all users upgrade to VMD 1.9.3. 2.1 Extend PyMOL with additional scripts.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
